|
EIF4A1
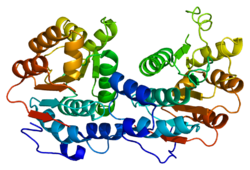
Eukaryotic initiation factor 4A-I (also known as eIF4A1 or DDX2A) is a 46 kDa cytosolic protein that, in humans, is encoded by the EIF4A1 gene, which is located on chromosome 17.[5][6][7] It is the most prevalent member of the eIF4A family of ATP-dependant RNA helicases, and plays a critical role in the initiation of cap-dependent eukaryotic protein translation as a component of the eIF4F translation initiation complex.[8] eIF4A1 unwinds the secondary structure of RNA within the 5'-UTR of mRNA, a critical step necessary for the recruitment of the 43S preinitiation complex, and thus the translation of protein in eukaryotes.[8] It was first characterized in 1982 by Grifo, et al., who purified it from rabbit reticulocyte lysate.[9] BackgroundThe regulation of the translation of mRNA transcripts into protein is one of the best ways that a cell can alter its response to its environment, as changes to the transcription of genes often takes considerably more time to be enacted. Protein translation can be broken into four phases: activation, initiation, elongation, and termination. Of these steps, initiation is the one for which cells have the most control. It is the rate limiting step of protein synthesis, controlled by a myriad of proteins known as the eukaryotic initiation factors, or eIFs. Relative abundance of these factors or their relative individual activities afford eukaryotic cells broad control over the rate of initiation and thus protein synthesis. eIFs are regulated under well known intracellular signalling pathways, such as the PI3K/AKT/mTOR pathway, however other biochemical layers of regulation, such as the complexity of RNA secondary structure in the 5′-UTR, are becoming evident with further research.[8] Examples of RNA Secondary Structure An example of an RNA hairpin, consisting of complementary Watson-Crick base pairings on the same strand of RNA. G-quadruplexes are more complex RNA secondary structures made up of stacks of guanine tetrads, which are each composed of four Hoogsteen hydrogen bonding guanines arranged in a square planar fashion. The eIF4A subfamily in mammals is made up of three paralogs, eIF4A1, eIF4A2, and eIF4A3.[10] eIF4A1 and eIF4A2 share 90% sequence similarity, and are both cytoplasmic proteins, while eIF4A3 is localized to the nucleus, and shares only 60% homology.[10] Historically, eIF4A1 and eIF4A2 were considered interchangeable, due to this being observed in in vitro experiments, but further investigation has shown that eIF4A1 is more prevalent in dividing cells while eIF4A2 is more abundant in non-dividing cells, and furthermore, more recent evidence suggests that they might have functionally distinct roles in vivo.[8][10] StructureeIF4A1 is a member of the DEAD box family of RNA helicases.[11] RNA helicases are enzymes that use the energy released from the hydrolysis of ATP to manipulate the secondary structure of RNA, and the DEAD box family is the largest family of RNA helicases.[11] The name "DEAD box" refers the key D-E-A-D amino acid sequence on motif II of the helicase that participates in nucleoside triphosphate binding (in the instance of eIF4A1, ATP). Other conserved motifs, shared by all eIF4A family proteins, are the Q, I, Ia, Ib, III, IV, V and VI motifs. Motifs Ia, Ib, IV and V bind RNA, motifs I, II, and III mediate RNA-dependent ATPase activity, and motif VI, is required for both RNA binding and ATP hydrolysis.[10]  The DEAD box family is marked by a structurally highly conserved helicase core consisting of two RecA-like domains joined by a flexible hinge region around which the protein can open and close upon hydrolysis of ATP.[13][10][14] The cleft formed between these two domains forms the ATP-binding pocket.[11] The RNA molecule binds opposite to this binding pocket, stretching across each of the domains.[11] This core is flanked by variable auxiliary domains, which confer the unique function of each RNA helicase to them partly by allowing for specific binding to accessory proteins.[11] FunctioneIF4A1 is an ATP-dependent RNA helicase,[15] however the exact nature of its dependence on ATP for its function is still debated.[10] Although after ATP binding, the subsequent hydrolysis induces conformational changes in eIF4A1, other DEAD-box RNA helicases have been shown to possess helicase activity in the presence of nonhydrolyzable analogues of ATP, suggesting that binding, and not hydrolysis, is the more important element in regulating activity.[10] eIF4A1 is a component of the eIF4F translation initiation complex, along with eIF4E, the 5'-terminal cap binding protein, and eIF4G, the scaffold protein that holds eIF4A and eIF4E together.[10] The eIF4F complex is often accompanied by the accessory proteins eIF4B and eIF4H, either of which can differentially enhance the activity of eIF4A1. After mRNA is transcribed from DNA and translocated to the cytoplasm, and the cytosolic PABP is bound to the Poly(A)-tail of the nascent mRNA, its 5'-cap will bind to eIF4E and PABP will bind to eIF4G.[8] eIF4A1 will then unwind the RNA secondary structure from 5' to 3' as the 43S PIC is recruited to the eIF4F complex.[8] The 43S PIC will scan the unwound mRNA from 5' to 3' as well, until it reaches the AUG start codon, whereupon the 60S ribosomal subunit will be recruited to begin the process of elongation.[8]  (B) eIF4A1 unwinding of mRNA secondary structure and recruitment of the 43S PIC. (C) 40S ribosomal subunit scanning the 5'-UTR of the mRNA transcript for a start codon. (D) Recruitment of the 60S ribosomal subunit and the beginning of elongation. RegulationThe transcription of eIF4A1 is driven by the transcription factor MYC.[8] On its own, the helicase activity of eIF4A1 is poor, however this feature imposes a practical restraint on eIF4A1, as nonspecific, "unintended," helicase activity in the cell would be detrimental to the function of certain endogenous, necessary RNA structures.[10] Its effectiveness considerably improves in the presence of eIF4B and eIF4H, binding partners that modulate its activity. When eIF4B binds to eIF4A1, the helicase activity of eIF4A1 is increased over 100-fold, but when eIF4H binds instead, the increase is not nearly as great, suggesting different relative concentrations of these accessory proteins may confer a further level of regulation of the efficiency of eIF4A1.[10] Conversely, eIF4A1 activity is suppressed when it is bound to PDCD4, a tumor suppressor itself modulated by mTOR and miR-21.[8] PCDC4 is typically localized to the nucleus in healthy cells, however, under carcinogenic conditions, it translocates to the nucleus and two separate eIF4A1 molecules will bind to it, inhibiting the ability of eIF4A1 to bind to RNA by locking the molecules into their inactive conformation, thereby preventing binding to eIF4G.[16][11] Role in DiseaseCancerTranslational dysregulation is a hallmark of malignant transformation of cancer cells. Cancer cells in growing tumors become "addicted" to heightened levels of protein translation, and particularly dependent on up-regulated translation of pro-oncogenic mRNAs. These pro-oncogenic mRNAs have characteristically longer 5'-UTRs with more complex secondary structures, and up-regulation of eIF4A1 has been implicated in several human cancers (See Table).[8][17][18] Given the general trend of eIF4A1 overexpression driving cancer, there is interest in developing inhibitors for the enzyme. Several natural compounds have been identified as candidate inhibitors for development, though they inhibit both eIF4A1 and eIF4A2 non-specifically.[8] These include hippuristanol, silvestrol and pateamine A, among others.[8] Silvestrol, in particular is a rocaglate derivative, and this class of compounds could be viable eIF4A inhibitors.[19] Putative eIF4A Inhibitors Hippuristanol Silvestrol Pateamine A
Viral InfectionsViruses rely on hijacking the cellular machinery of the cells they infect to create their own viral proteins and allow them to continue infecting new cells. Their ability to manipulate eIFs like eIF4A1, therefore, considerably impacts their virulence. For instance, cytomegalovirus relies on eIF4A to drive its protein synthesis. The viral protein pUL69 stabilizes the formation of eIF4F, through binding to eIF4A, a process by which eIF4E is prevented from dissociating from the eIF4F complex.[14] eIF4E, thus, is no longer able to be sequestered by its negative regulator, 4EBP.[14] Furthermore, cytomegalovirus stimulates the synthesis of all elements of the eIF4F complex in order to drive protein synthesis.[14] Other viruses, like Cotesia plutellae bracovirus (CpBV), that favor cap-independent translation, will take advantage of eIF4A1 in the reverse context, by sequestering eIF4A1 away from the eIF4F complex with viral binding partners, in this case a protein called CpBV15β, thus inhibiting endogenous cap-dependent mRNA translation and favoring viral protein translation.[14] The compounds mentioned in the above section about cancer, hippuristanol, silvestrol, pateamine A, rocaglate derivatives, etc., could also be applied as putative viral inhibitors.[8][19] References
Further reading
External linkspUL69 of cytomegalovirus (UniProt) CpBV15β of Cotesia plutellae bracovirus (UniProt) |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||